Bacterial Genetics and Genomics book Discussion Topic: Chapter 3, question 13
One of the concepts that is discussed in Bacterial Genetics and Genomics is the difference between essential genes and accessory genes. The essential genes are those that are required for the bacteria to live and the accessory genes allow it to do something else that might be beneficial, but not essential. This is the difference between having all of the genes needed for cellular division and having the genes needed for a bacterial capsule. Being able to divide is an essential function and without it, the bacteria are not viable. The bacterial capsule may enable the bacteria to avoid the human immune system thus boosting its survival, but it is not essential. In this comparison, those that are essential and those that are accessory are determined within the bacteria itself, not by comparing between bacteria – even those of the same species.
Another concept discussed is the difference between core genes and accessory genes. In this case, we are comparing the genes present between two different bacteria of the same species. The genes that are in common between the bacteria are core. Those that are not are accessory. This is often represented with a Venn diagram, showing all of the genes that are shared between two or more bacterial strains – the core genes – and those that are not and are accessory genes. Everything together is the pan-genome.
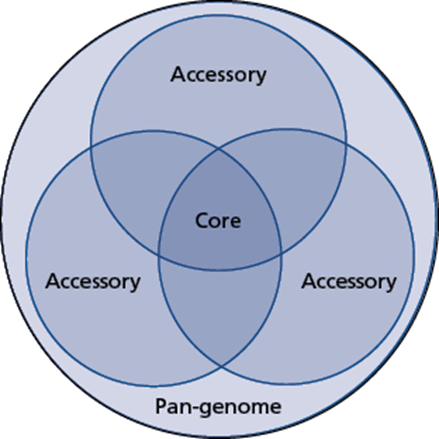
It is interesting that both of these concepts use the term accessory gene – something that could easily confuse students and researchers when discussing the nature of these genes. The point being that the genes in question are not essential to the function of the bacterial cells and quite often are not present in the genomes of all examples of that species.
In looking into this topic in the research literature, I came across a paper in PNAS (Proceedings of the National Academy of Sciences of the United States of America) by Bradley Poulsen and colleagues in Massachusetts, published in 2019. This article begins with a quite frank overview of what was believed to have been the promise of bacterial genome sequencing when it comes to the development of new antimicrobials. As they describe it, the revolution that was set to transform antibiotic discovery through the ability to identify essential genes and target them for new drugs, made possible from the first bacterial genome sequence in 1995, has just simply failed to materialize.
There are, of course, a number of problems that have arisen for which genomics is not entirely to blame. As noted in the PNAS paper, generally the bacterial membrane and efflux pump systems need to be overcome for any new drugs to be able to reach their targets. This is a strength of our bacterial foes, not a weakness of our genomic methods. Finding antimicrobials in some cases starts with screens of chemical libraries and improvements on these are often needed, which is again not a shortcoming of genomics. Lastly, there have been some successes in identification of new antibiotics, but often these are not broad-spectrum agents and therefore their development is abandoned or not picked up by pharmaceutical companies for further development and clinical trials due to the lack of profitability. This is again, not the fault of genomics.
However, the definition of essential genes does fall within genomics. In some cases, targets have been pursued that have not actually been essential, at least not in all species or in all strains of a species. This results in inhibitors having been developed to bacterial targets that are only effective against a sub-population of the pathogen, which is pretty much useless in a clinical setting.
In addition, essential genes can be essential for a particular set of circumstances. They can be essential depending on how the bacteria are being grown, for example, but if the bacteria are grown in different conditions those genes are no longer needed and would have otherwise been classified as accessory genes. These are ‘conditional essential genes’ – ones that are essential only under certain conditions. The bacteria may not encounter that condition or may be able to avoid being in that condition within the patient and therefore could avoid being cleared by the antibiotic developed to target the essential gene by not needing to use the gene and staying somewhere else in the body. It is quite common for genes to be essential in laboratory media, which is an artificial environment, but entirely different sets of genes to be needed for the various environments encountered in the body, like the gut, the blood, the mucosal surfaces, and inside cells.
Poulsen et al. proposed in their 2019 PNAS paper that antibiotic drug discovery could be improved through focusing on core essential genes – those that are shared amongst all of the sequenced genomes of a species AND that could be demonstrated to be essential in a range of different conditions that mimic the human body, not just one. In their proof-of-concept investigation, they chose to look at Pseudomonas aeruginosa due to the urgent need for novel antibiotics.
To find essential genes, they used transposon insertion sequencing. This is known as Tn-Seq, TIS, INseq, HITS, and TraDIS. Two P. aeruginosa laboratory strains were mutated in this way, strain PA14 and PAO1. Eight other strains were also investigated under five growth conditions and a statistical method called Finding Tn-Seq Essential genes (FiTnEss) was used to identify the core essential genes from the data collected.
In the end, the conclusion of the authors is that it is not genomics that has failed us in providing answers that we need for developing new antibiotics to combat the growing threat of antimicrobial resistant infections. It is rather our own inability to use genomics to its best advantage and to design experiments to best identify the genes that need to be targeted by novel antimicrobials. Through the advances in genomic technologies, it is now possible to rapidly and inexpensively sequence and re-sequence bacterial genomes, which was not possible in the 1990’s, 2000’s, and even 2010’s. At the start of the 2020’s, we are able to use the advances of genomics to do the types of experiments we have wanted to do, but have often been out of reach due to the expense and scope that would have been out of reach previously.
Therefore, the quest for core essential genes is has only just begun.
