Bacterial Genetics and Genomics book Discussion Topic: Chapter 11, question 14
Although I had planned to continue to work through the Discussion Topics from Bacterial Genetics and Genomics that are all related to the investigations in bacterial genetics and genomics using bioinformatics tools can be conducted outside of the lab, October here in the UK includes Biology Week. In addition to my research and teaching at the University, I also like to do outreach activities and public engagement in science, which includes going to schools to talk about biology, my career as a woman in science, and the cool things that we can do with genomics. One of the stories that the students and teachers find particularly fascinating is one that I was asked to deliver for Biology Week this year, but remotely, due to COVID-19. So, for this month’s blog, I have combined by regular blog with a video that I have produced for Biology Week about the outbreak of cholera in Haiti and how genomics helped to solve the mystery of its origin.
Bacterial Genomics – Cholera in Haiti
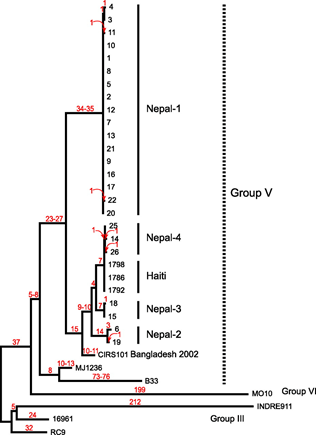
In January 2010, an earthquake struck Haiti causing widespread devastation. The international aid community responded, with the UN coordinating sending troops to help rebuild needed infrastructure and help the population. However, in October 2010, there was an outbreak of cholera that was traced back to the UN aid worker station and unsanitary conditions of waste disposal there. Ultimately, genome sequencing of the Vibro cholerae bacteria showed that it was nearly identical to bacteria from a recent outbreak in Nepal (doi: 10.1128/mBio.00157-11), as shown in the phylogenetic tree figure below. This genomic information links the outbreak to the arrival in October 2010 of UN aid workers from Nepal. Although none of these people who had come to help had symptoms, this emphasizes the role that asymptomatic carriage has in the spread of infections during an outbreak.
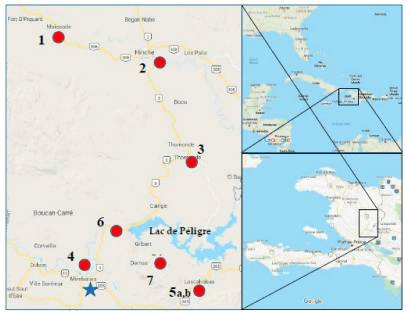
Using metagenomics, the sequencing of environmental samples of DNA, researchers investigated the prevalence of cholera in Haiti in the years following the initial outbreak. Monika A. Roy and co-authors published in 2018 their metagenomic approach to evaluating surface water quality in Haiti (doi: 10.3390/ijerph15102211). In their publication, they note that since the earthquake in January 2010 and the subsequent cholera outbreak in October 2010, there had been over 9,000 deaths from cholera in Haiti.
During the term of their investigation, cholera cases had declined, but were still occurring and one of the issues was the inability to predict where new cases might occur due to gaps in surveillance. Through use of metagenomics to monitor groundwater it was hoped this might identify sources of potential cholera contamination.
The cost of using genomics for metagenomics monitoring of environmental samples is mentioned in the Introduction of the paper by Roy et al., 2018. They mention the availability of handheld MinION devices, such as those discussed in my previous blog entry, but cite that the regents to run one flowcell cost $1000. They concede that if samples are multiplexed, mixed together with molecular barcodes to differentiate them so more samples can be run at once, then the per sample cost can be reduced to $80 per sample.
Water samples were collected in 2017 as a pilot investigation and then again from specific locations in triplicate in 2018. The water was filtered using 0.22 μm filters. This is the same size filter we would use in the lab to make solutions that cannot be autoclaved, but which we need to be sterile. Therefore, bacteria in the water would be trapped onto the filters and the DNA from these was later extracted in the lab using a QIAGEN kit. This DNA is a mixture of DNA from the environment, thus being a metagenomic sample, which was sequenced using an Ion Torrent sequencer. The data was processed using CosmosID to gain the results that were interpreted by the team.
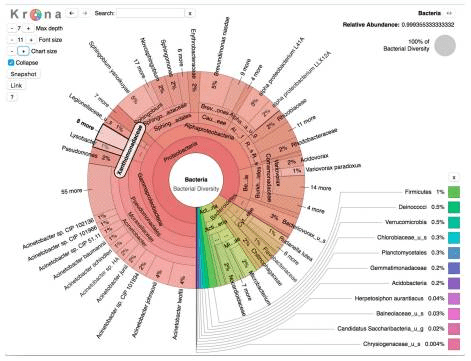
Another issue is the bioinformatics, but again the authors acknowledge that there are improvements here in making workflows easier to use and better optimized. Using the popular Krona visualization tool for metagenomics analysis, the diversity of organisms revealed is presented. They noted that V. cholerae was present in most of the replicates of water samples, although at varying abundance. The researchers are careful in their experimental design and note that V. cholerae is not always pathogenic; this is also an environmental bacterial species and when investigating metagenomics, markers must be selected to identify pathogenic versus non-toxigenic strains. The Haitian strain HE-45 was detected and samples with the virulence gene V. cholerae intI1 were identified. Cholera toxin converting phage was detected in one sample from a source not known to be used as drinking water, however these places are used for bathing, washing clothes, and for other household activities that could result in accidental ingestion of contaminated water. In addition to the cholera toxin gene, sequences for Shiga toxin were also detected. A distinct advantage of a metagenomics approach is the identification of sequences that were perhaps not part of an original hypothesis; if the initial concern had been about cholera and the investigation had just looked for cholera, these other sequences of concern would have been missed. Shiga toxin is made by EHEC E. coli such as O157:H7 and can cause life threatening disease.
The conclusion of the metagenomics study was that no V. cholera O1 or O139 strains were detected, which was consistent with the decline in cholera cases, however the presence of cholera toxin genes and Shiga toxin genes in the water was concerning. These could indicate potential risks for the populations using this water.
The last case of cholera in Haiti was seen in January 2019. Control of the outbreak has taken a tremendous effort in re-establishing clean water and sanitation that was destroyed by the hurricane and improving upon the infrastructures of Haiti, as well as implementing other control measures such as vaccination and rapid tracing and treating systems. In April 2019, there were concerns that the apparent eradication of cholera in Haiti should not be taken as a sign that the nation could cease being vigilant. High standards in potable water, sanitation, and healthcare must still be emphasized as of fundamental importance for preventing outbreaks across the globe.
